Want to share your content on R-bloggers? click here if you have a blog, or here if you don’t.

The R Consortium Submission Working Group has now successfully made two pilot submissions to the FDA. All the submissions done by the group are focused on improving practices for R-based clinical trial regulatory submissions. Now, the R submission Working Groups, in collaboration with Appsilon and Posit, are exploring new technologies such as Containers and WebAssembly. In this article, we dive into the details of this exploration.
Table of Contents
- How Everything Started
- What’s next?
- The Journey with WebAssembly and Containers
- The Pilot 4 Shiny App Up and Running on webR!
- webR Shiny App
- Next Steps
How Everything Started
Pilot 1
This pilot was initially submitted on November 22, 2021. This submission was the first publicly available R-based submission to the FDA. This was a test submission that aimed to explore the submission of an R package to the FDA following the eCTD specifications. The submission included an R package, R scripts for analysis, R-based analysis data reviewed guide (ADRG), and other important components. The final response letter from the FDA was received on March 14, 2022.
Pilot 2
This was one of the first submission packages containing a Shiny application. The main goal of this pilot was to test the submission of an R-based Shiny application bundled into a submission package and transfer it successfully to FDA reviewers. The submitted application was built using the datasets and analyses that were used for the R Submission Pilot 1. The deployed version of this application is available on this site. Alternatively, a Rhino-based version of the application can be found here.
The final response letter from the FDA was reviewed on September 27, 2023.

What’s next?
Pilot 3
This pilot was successfully submitted to the FDA on Aug 28, 2023. This was the first publicly available R submission that included R scripts to produce ADaM datasets and TLFs. Both the ADaMs (SDTM .xpt sources from the CDISC Pilot study) and the TLFs (ADaMs .xpt sourced from the ADaMs generated in R by the Pilot 3 team) were created using R. The next step for this pilot is to await FDA’s review and approval, which may take several months to complete.
Pilot 4
This pilot aims to explore using technologies such as containers and WebAssembly software to package a Shiny application into a self-contained unit, streamlining the transfer and execution process for enhanced efficiency.
This pilot is expected to be divided into two parallel submissions:
(a) will investigate WebAssembly and
(b) will investigate containers.
The Journey with WebAssembly and Containers
Our team at Appsilon teamed up with the dynamic Pilot 4 crew to explore WebAssembly technology and containers. George Stagg and Winston Chang also joined the working group to discuss the web-assembly portion of Pilot 4. This partnership brought together our engineering prowess to contribute to these tools, injecting fresh perspectives into the ongoing pilot project.
Some of the outcomes of the collaboration:
- We were able to set up a robust container environment for this pilot project.
- We aided the progress made on the use of both experimental technologies: containers and WebAssembly.
- We developed a working prototype submission using Podman container technology.
- We developed a working early-stage prototype for wrapping a small Shiny application using WebAssembly.
WebAssembly

webr
WebAssembly allows languages like R to be executed at near-native speed directly within web browsers, providing users with the ability to run R code without having R installed locally. WebR is essentially the R programming language adapted to run in a web browser environment using WebAssembly. This project is under active development.
The Pilot 4 Shiny App Up and Running on webR!
The deployed example of the Shiny app running on webR is available here. Check out the video of the application running below.
Below is the dockerfile (recipe) for the container:
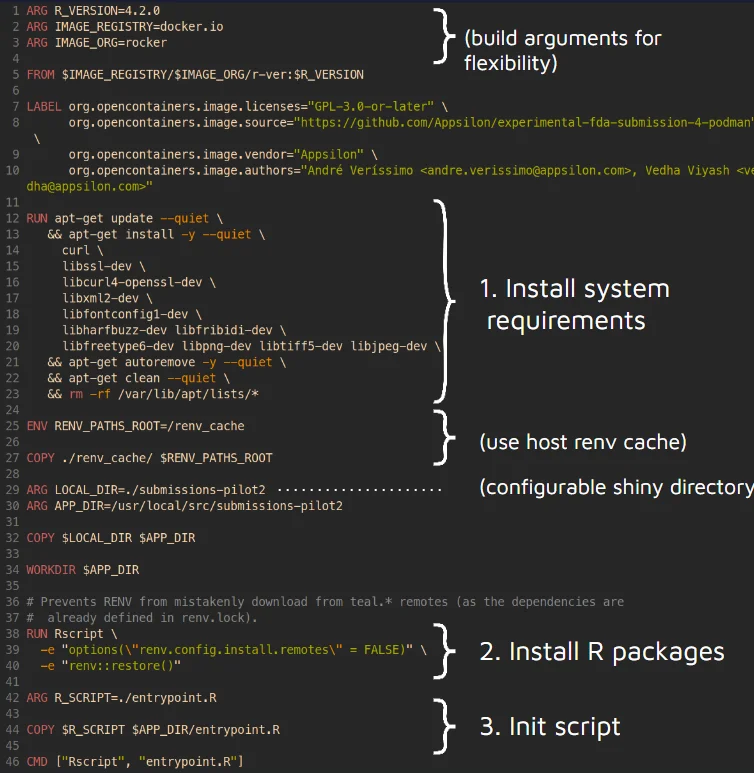
Next Steps
The next steps are waiting for the review of Pilot 3 by the FDA and to submit the two sections that explore the new technologies to regulatory authorities. Thanks to the collaboration between the R Submission Working Group and other institutions, there is already a working prototype of a {teal}-like Shiny application running on webR and further exploration with Podman is underway.
- Submission to FDA
- Rhino Compatibility

Appsilon is working on the side with Rhino compatibility; eventually, we might be able to just add this framework into the Pilot 4 application.
- {teal}This and other packages might be on CRAN soon. We could incorporate them after that, replacing the shims created for this version.
- Boot TimeWe need to improve boot time (remove dependencies and keep working on webR).
This article was co-authored by Pedro Silva, André Veríssimo, Vedha Viyash & Tymoteusz Makowski
This post was originally published on https://pharmaverse.github.io/.
The post appeared first on appsilon.com/blog/.
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you’re looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don’t.
Continue reading: Testing Containers and WebAssembly in Submissions to the FDA
Analyzing the Cutting-Edge Work of The R Consortium Submission Working Group
The R Consortium Submission Working Group has been pushing boundaries in the field of R-based clinical trial regulatory submissions. Our main discussion points will be centered around its innovation in executing four pilot submissions to the FDA and its exploration of new technologies such as Containers and WebAssembly in collaboration with Appsilon and Posit.
Improving Clinical Trial Regulatory Submissions
The R Consortium Submission Working Group has achieved considerable progress with two successful pilot submissions to the FDA. These submissions reveal an exploration of novel practices for producing R-based clinical trial regulatory submissions. Examples include the submission of an R package following eCTD specifications and the introduction of a Shiny application within a submission package.
New Technologies on the Horizon
Looking ahead, the R Submission Working Groups are venturing into promising technologies like Containers and WebAssembly. Through their collaboration with Appsilon and Posit, these technologies hold potential for streamlining and enhancing the efficiency of the submission process.
Potential Long-Term Implications
The successful use of these technologies could lay a foundation for their wider adoption in other areas. Not only could this streamline regulations and submissions in the clinical trials landscape but it could also have broader implications for how data and software applications are handled in other sectors.
Boosting Efficiency with Container and WebAssembly Technology
Pilot 4 holds a special interest as it is set to explore both Containers and WebAssembly. Containers allow software to run reliably when moved from one computing environment to another, enhancing resource efficiency. On the other hand, WebAssembly permits code to execute at near-native speed within web browsers, providing users the flexibility to run R code without a local installation. A successful show of these technologies could lead to more accessible and efficient R-based applications and submissions.
Actionable Advice
- Organizations looking to improve their submission process should closely monitor the successes of the R Consortium Submission Working Group’s pilots.
- Consider adopting new technologies like Containers and WebAssembly to streamline your R-based application development and deployment process.
- To reduce dependencies and enhance execution speed, consider wrapping smaller applications with WebAssembly. Transitioning to a containerized application environment can aid in creating reproducible and portable software deployments.
- Companies should also keep an eye out for emerging IG packages like {teal} that could perform useful tasks such as analyzing clinical trial data.
In conclusion, the work of the R Consortium Submission Working Group presents valuable lessons on improving R-based clinical trial regulatory submissions and highlights the potential benefits of embracing new technologies. Their continuing explorations stand to offer insights for improving efficiency, accessibility, and versatility in R-code execution.
